Scientists at Rice University’s Center for Theoretical Biological Physics (CTBP) are part of a study to develop an inhalable COVID-19 vaccine.

The project led by Rutgers University and CTBP scientists at Rice and Northeastern University has produced two vaccine strategies. Both are scalable and adaptable and can be transported and stored at room temperature.
One strategy employs modified bacteriophage particles that can be inhaled to deliver protection via the lungs to the immune system. The other delivers injectable adeno-associated virus-phage particles that directly encode protection against the virus in immune cells.
Both approaches triggered strong production of antibodies in rodents.
The study appears in the Proceedings of the National Academy of Sciences.
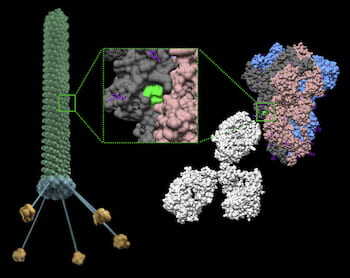
Rice physicist José Onuchic, a co-principal investigator on the project, and his team worked on the first strategy. CTBP scientists simulated several epitopes, the part of antigen molecules that bind to specific biological targets. These can be placed on the surface of bacteriophage (aka phage) particles, viruses that infect bacteria but are safe for humans and have been used to treat bacterial infections for nearly a century.
The phage particles simulated at CTBP and developed at Rutgers are engineered with an epitope from the SARS-CoV-2 spike protein, along with a small ligand peptide that helps the phage cross from the lungs into the recipient’s bloodstream. Once there, they essentially teach the immune system to guard against COVID-19.
An advantage to a phage-enabled vaccine, Onuchic said, is that one spike protein can carry a multitude of epitopes, and they can be easily customized to protect against COVID-19 variants. “The protection derived from some of these epitopes may be destroyed in a variant, but the remaining ones will continue to offer protection,” he said.
The Rice team’s initial task was to see which of a small set of epitopes displayed by SARS-CoV-2 would best serve the purpose.
“We wanted to find fragments that mimic the spike structure, so they can be used to teach the immune system to recognize the virus,” said co-author Paul Whitford, a CTBP senior scientist and an associate professor of physics at Northeastern.
“Our work demonstrated the epitopes that show the smallest deviation from the original structure when put onto the surface of the phage are the most efficient in terms of immune response,” Onuchic said. “This is because their structures are mostly conserved when they are taken out of the protein environment. It appears that not only sequence but also structural conservation is needed for success.
“This also gives us the possibility of designing new epitopes, and now we have a methodology to look for them computationally,” he said.
In this study, CTBP scientists, including co-authors Esteban Dodero-Rojas, a Rice graduate student, and Vinicius Contessoto, a former postdoctoral researcher at Rice and currently a CTBP affiliate, analyzed five epitopes suggested by Rutgers scientists and found that one of them was best able to retain its structure when transferred to a phage.
In experiments with rodents, that one, which they called Epitope 4, delivered the best immune response by nearly a factor of 10 over the other candidates.
In the next step, the researchers will expand the search for more and better epitopes.
“We will try to computationally screen the entire surface of the spike protein to find new epitopes,” Onuchic said. “If you just do this by trial and error, by brute force, it’s going to be too expensive or too long a process. By computational screening, we can come up with a short list of new candidates.”
“This paper proves the concept that, as of the last five or six years, it has become easier and cheaper to test vaccine constructs on the computer prior to experiments,” Whitford added. “Before, one could typically test just a handful of possibilities, but now we’re ramping up to simulate hundreds or thousands of candidates.”
A single vaccine that protects against multiple COVID-19 variants and can be transported without the “cold chain” required for current vaccines will be key to finally curtail the worldwide epidemic, said Renata Pasqualini, who co-led the study.
“The structure and strength of the local health care system is a key consideration,” said Pasqualini, a professor and founding chief of the Division of Cancer Biology in the Department of Radiation Oncology at Rutgers New Jersey Medical School. “As such the global COVID-19 pandemic has raised awareness of public health inequity and the need for a rapid and accessible immunization process. In this context, the favorable biological attributes of phage particles might represent a potentially practical yet affordable alternative.”
Onuchic said the study serves as a powerful example of how theory and experimentation should work together. “You’re going to see a lot of papers going forward that are not theoretical or experimental, but a synergistic combination of both,” he said.
“I was amazed at how rapidly yet efficiently we assembled and integrated a multidisciplinary team of molecular, cell and structural biologists to design and test a preclinical candidate vaccine,” said co-lead author Wadih Arap, director of the Rutgers Cancer Institute of New Jersey at University Hospital Newark.
“Ongoing and planned studies will hopefully confirm that our first prototype is indeed neutralizing and leads to an Investigational New Drug application to the FDA,” he said. “In the meantime, the platform technology reported in this work will serve to respond promptly to emerging more virulent variants.”
Additional co-authors are Daniela Staquicini, Fenny Tang, Christopher Markosian, Deodate Davis, Paul O’Brien, Nazia Habib, Tracey Smith, Edmund Lattime, Steven Libutti and Stephen Burley of the Rutgers Cancer Institute of New Jersey; Virginia Yao of PhageNova Bio; Fernanda Staquicini of MBrace Therapeutics; Natalie Bruiners and Maria Gennaro of the Rutgers New Jersey Medical School; and Richard Sidman of Harvard Medical School.
Onuchic is the Harry C. and Olga K. Wiess Chair of Physics and a professor of physics and astronomy, chemistry and biosciences and CTBP co-director.
The Rutgers Cancer Institute of New Jersey, National Science Foundation, Department of Energy, National Institutes of Health, Northeastern University, a Gillson Longenbaugh Foundation grant and the Welch Foundation supported the research.